“Between 30% to 80% of the data analysis task is spent on cleaning and understanding the data.” (Dasu & Johnson, 2003)
In this section, we use the starwars dataset from the dplyr package to review the main tools for data manipulation in R.
library(dplyr)
# We will refer to the starwars dataset as sw
sw = starwars
# Drop the last three columns to simplify the examples.
# These columns are lists.
sw[, c("films", "vehicles", "starships")] = NULL
The base package
- In this section, we manipulate datasets using the functions that are loaded by default in R.
Summarizing data
Basic functions
- Summarizing data (John Hopkins/Coursera)
- We will inspect:
- the dataset dimensions with
dim() - the first six rows with
head() - the last six rows with
tail()
- the dataset dimensions with
dim(sw) # dataset size (rows x columns)
head(sw) # first six rows
tail(sw) # last six rows
- With
str(), we can inspect the structure of the dataset:- all variables (columns)
- the class of each variable
- a sample of the observations
str(sw)
- To generate a summary for all variables, we can use
summary(). For numeric variables, the function reports quantiles, the mean, and the number of missing values.
summary(sw)
- For logical, text, or categorical variables,
summary()is usually less informative. In those cases, it is often useful to build a frequency table withtable()and then turn it into percentages withprop.table(table()).
table(sw$hair_color) # counts
prop.table(table(sw$hair_color)) # percentages
- We can also build a cross-tabulation by including more than one variable inside
table():
table(sw$hair_color, sw$gender)
The apply family
The apply family provides compact ways to repeat operations:
apply(): applies a function to the rows or columns of a matrix or arraylapply(): applies a function to each element of a listsapply(): similar tolapply(), but tries to simplify the output
The apply() function
- Loop functions - apply (John Hopkins/Coursera)
apply()is commonly used to evaluate a function over the rows or columns of a matrix.- It is not necessarily faster than writing an explicit loop, but it is concise and readable.
apply(X, MARGIN, FUN, ...)
X: an array, including a matrix.
MARGIN: a vector giving the dimensions over which the function will be applied.
FUN: the function to be applied.
...: optional arguments passed to FUN.
x = matrix(1:20, 5, 4)
x
apply(x, 1, mean) # row means
apply(x, 2, mean) # column means
- There are built-in functions that reproduce these operations directly:
rowSums = apply(x, 1, sum)rowMeans = apply(x, 1, mean)colSums = apply(x, 2, sum)colMeans = apply(x, 2, mean)
The lapply() function
- Loop functions - lapply (John Hopkins/Coursera)
lapply()takes three main inputs: a list, a function, and optional additional arguments.
lapply(X, FUN, ...)
X: a vector (atomic or list) or an expression object.
FUN: the function to be applied to each element of X.
...: optional arguments passed to FUN.
lapply()applies a function to each element of the list.- A data frame is a special kind of list whose elements are its columns.
lapply(sw, mean, na.rm = TRUE)
Notice that non-numeric variables return
NA, and the third argument above is an argument ofmean().We can also inspect the unique values of each variable by combining
lapply()withunique():
lapply(sw, unique)
- A particularly useful application is counting the number of missing values in each column. Because
is.na(sw)returns a matrix, we can combine it withapply():
head(is.na(sw)) # first rows after applying is.na()
class(is.na(sw)) # object type
apply(is.na(sw), 2, sum) # sum TRUE/FALSE by column
The sapply() function
sapply()is similar tolapply(), but it tries to simplify the result whenever possible.- If each list element returns an object of the same length,
sapply()often returns a vector or matrix instead of a list.
sapply(sw, mean, na.rm = TRUE)
Filtering rows
# logical vector for blond hair
sw$hair_color == "blond"
# extract rows with blond hair
sw[sw$hair_color == "blond", ]
- We can use logical expressions to filter observations. For example, suppose we want characters with blond hair and non-missing
hair_color:
sw[sw$hair_color == "blond" & !is.na(sw$hair_color), ]
- To avoid repeating the dataset name before each variable, we can use
with():
with(sw,
sw[hair_color == "blond" & !is.na(hair_color), ]
)
- We could also extract observations with blond or white hair:
# extract rows where hair is blond or white
sw[sw$hair_color == "blond" | sw$hair_color == "white", ]
- We can also check whether values belong to a given vector with
%in%, which is equivalent to applying==to several values at once:
sw$hair_color %in% c("blond", "white") # TRUE/FALSE for blond or white hair
sw = sw[sw$hair_color %in% c("blond", "white"), ]
head(sw)
Reordering rows
- Subsetting and sorting (John Hopkins/Coursera)
- We can use
sort()to order a vector in ascending order (default) or descending order.
sort(sw$height) # ascending order
sort(sw$height, decreasing = TRUE) # descending order
sort()is not appropriate for reordering a full data frame, because it returns only a vector.- To reorder data frames, we use
order(), which returns the row indices from the smallest values to the largest.
order(sw$height) # row indices in ascending order of height
sw = sw[order(sw$height), ] # reorder the data frame
head(sw)
Selecting columns
- We can select columns with
[,]by passing a vector of names or indices to keep.
# Select the first three columns
head(sw[, c("name", "height", "mass")])
# Select the first six columns
sw = sw[, 1:6]
head(sw)
Renaming columns
- We can rename variables with
names()orcolnames()by assigning a new vector of names.
names(sw) # current variable names
# Remove underscores
names(sw)[4] = "haircolor" # change one name
names(sw)[5:6] = c("skincolor", "eyecolor") # change two names
names(sw)
Modifying columns
- To modify a variable, we can assign a vector of the same length using
$.
sw$height = sw$height / 100 # convert cm to meters
- If we assign a scalar, R repeats it for all observations:
sw$const = 1
- We can also create new variables:
sw$BMI = sw$mass / sw$height^2 # body mass index
head(sw)
abs(sw$BMI[1:4]) # absolute value
sqrt(sw$BMI[1:4]) # square root
ceiling(sw$BMI[1:4]) # round up
floor(sw$BMI[1:4]) # round down
round(sw$BMI[1:4], digits = 1) # one decimal place
cos(sw$BMI[1:4]) # cosine
sin(sw$BMI[1:4]) # sine
log(sw$BMI[1:4]) # natural logarithm
log10(sw$BMI[1:4]) # base-10 logarithm
exp(sw$BMI[1:4]) # exponential
Merging datasets
A merge matches cases, that is, rows or observations, across two datasets using one or more key variables:
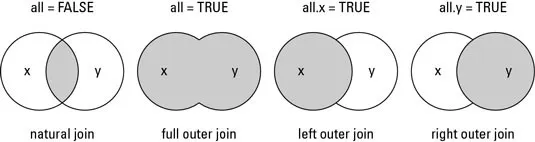
- We use the
merge()function for that purpose:
merge(x, y, by = intersect(names(x), names(y)),
by.x = by, by.y = by, all = FALSE, all.x = all, all.y = all,
suffixes = c(".x", ".y"), ...)
x, y: data frames, or objects that can be coerced into data frames.
by, by.x, by.y: columns used for merging.
all: shorthand for setting both all.x and all.y.
all.x: keep all rows from x.
all.y: keep all rows from y.
suffixes: suffixes used when non-key variables share the same name.
- Let us create two datasets where some individuals appear in one dataset but not in the other:
# Extract smaller data frames from the original starwars dataset
bd1 = starwars[1:6, c(1, 3, 11)]
bd1
bd2 = starwars[c(2, 4, 7:10), c(1:2, 6)]
bd2
- There are 12 unique characters across the two datasets, but only
"C-3PO"and"Darth Vader"appear in both. - To check which columns share the same name, we can combine
intersect()withnames()orcolnames():
intersect(names(bd1), names(bd2))
- If we do not specify the key variable,
merge()uses all columns with matching names.
Inner join
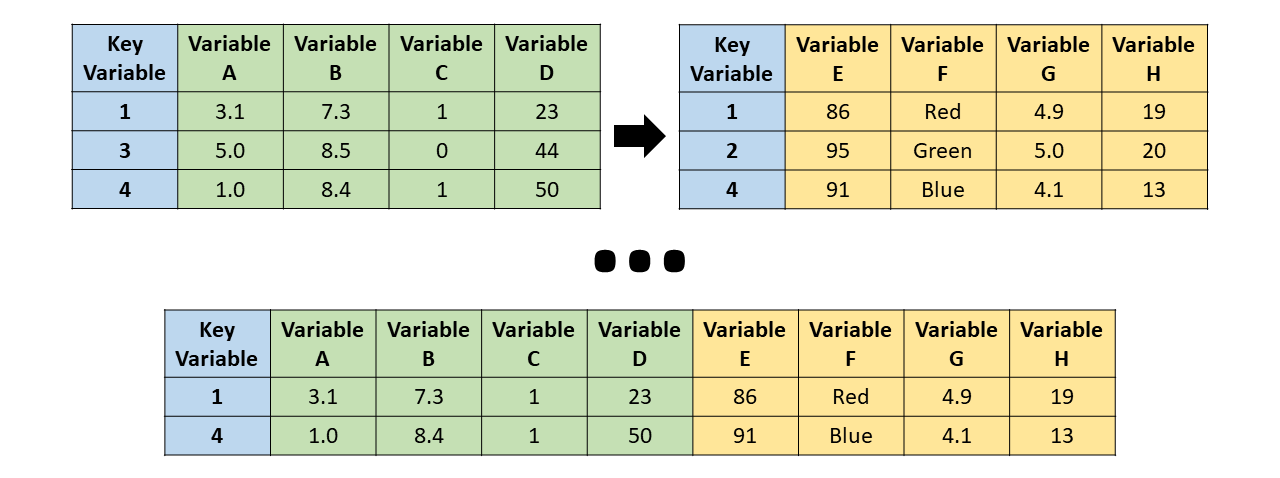
merge(bd1, bd2, all = FALSE)
Full join
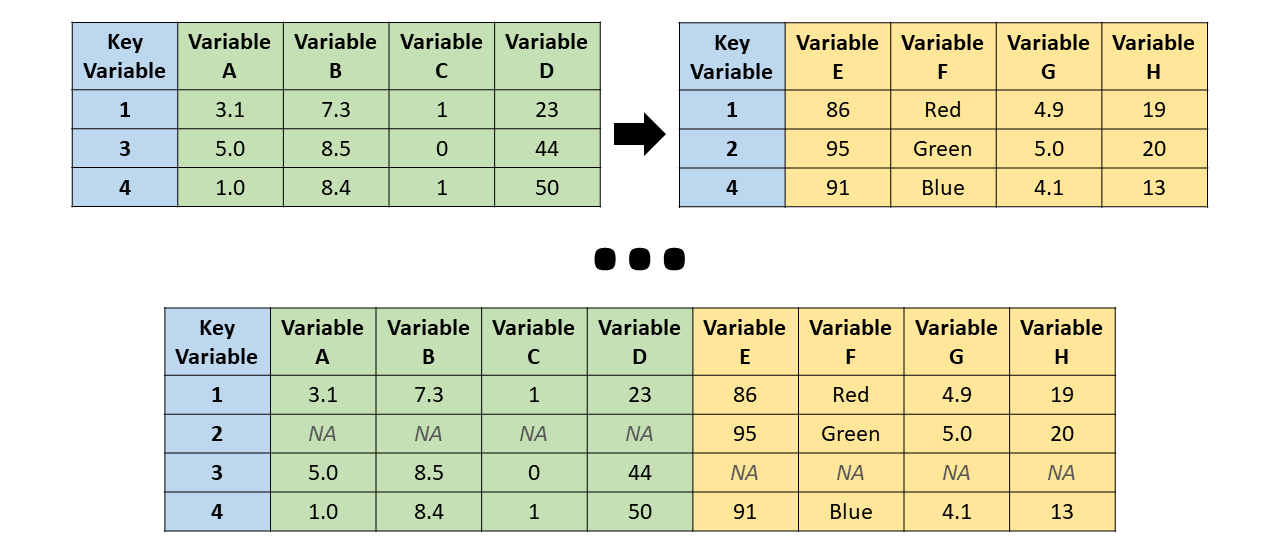
merge(bd1, bd2, all = TRUE)
Left join
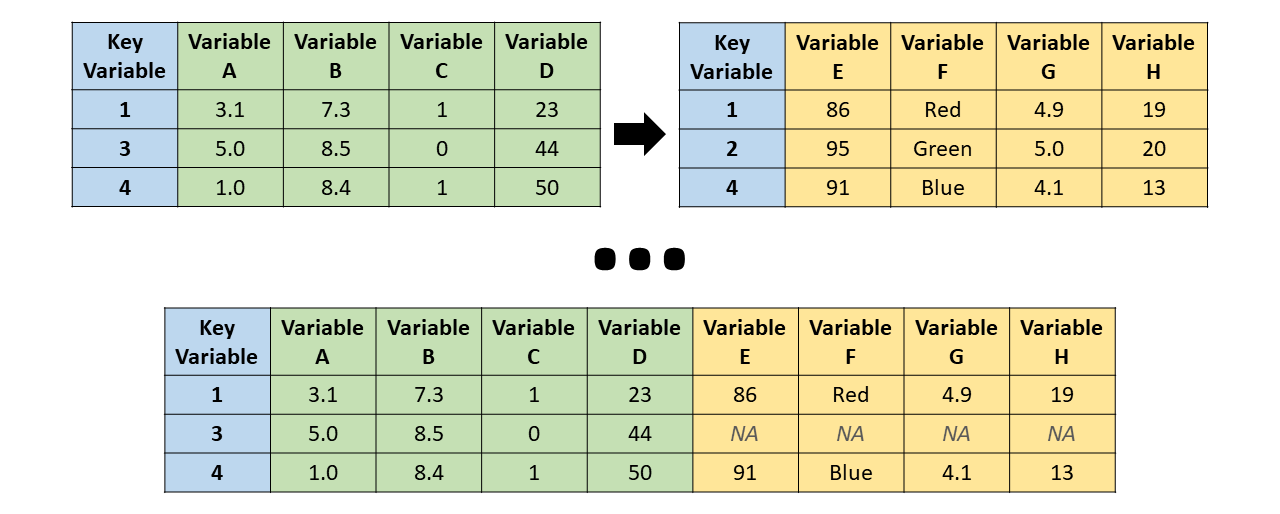
merge(bd1, bd2, all.x = TRUE)
Right join
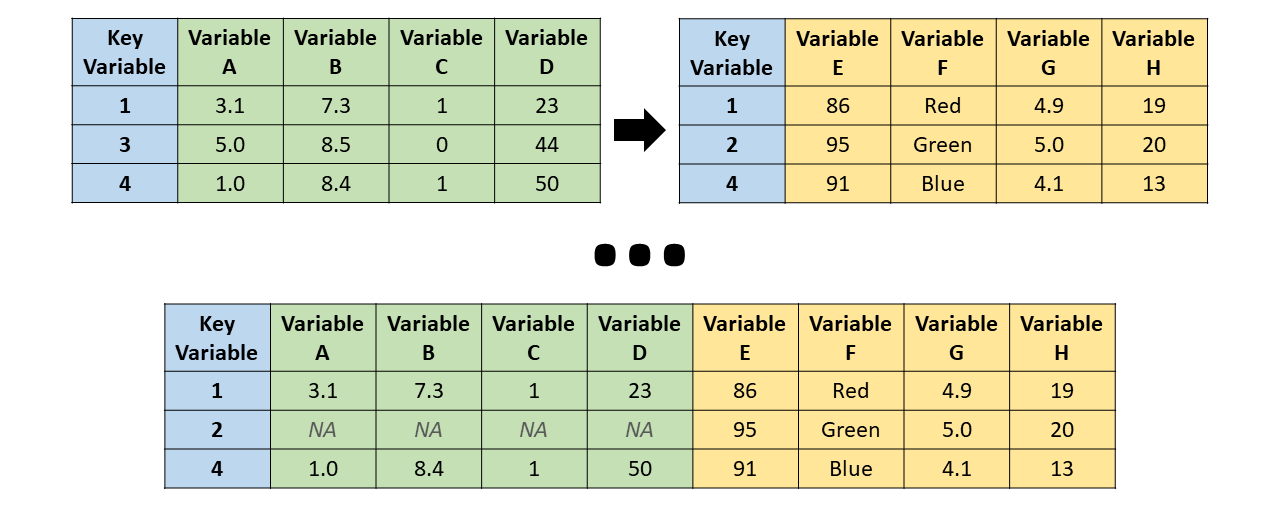
merge(bd1, bd2, all.y = TRUE)
The dplyr package
- Vignette - Introduction to dplyr
- The
dplyrpackage makes data manipulation easier through simple and computationally efficient functions. - These functions can be grouped into three categories:
- Columns:
select(): selects or drops data-frame columnsrename(): changes column namesmutate(): creates or modifies column values
- Rows:
filter(): selects rows according to column valuesarrange(): reorders the rows
- Groups of rows:
summarise(): collapses each group into a single rowgroup_by(): creates grouped data
- Columns:
- In this subsection, we continue using the Star Wars dataset.
- You will notice that, when these functions are applied, the data frame is turned into a tibble, which is a more efficient format for tabular data but works very similarly to a standard data frame.
library("dplyr") # load package
head(starwars) # inspect the first rows of the dataset included in the package
Filter rows with filter()
- Selects a subset of rows from a data frame.
filter(.data, ...)
.data: a data frame or tibble.
...: logical expressions defined in terms of the variables in .data.
starwars1 = filter(starwars, species == "Human", height >= 100)
starwars1
# Equivalent to:
starwars[starwars$species == "Human" & starwars$height >= 100, ]
Reorder rows with arrange()
- Reorders the rows according to one or more column names.
arrange(.data, ..., .by_group = FALSE)
.data: a data frame or tibble.
...: variables, or functions of variables. Use desc() for descending order.
- If more than one variable is provided, the data are first ordered by the first variable and ties are broken using the second.
starwars2 = arrange(starwars1, height, desc(mass))
starwars2
Select columns with select()
- Selects the columns of interest.
select(.data, ...)
...: variables in a data frame.
- We can list the variables we want to keep, select ranges with
var1:var2, or drop variables with-var.
starwars3 = select(starwars2, name:eye_color, sex:species)
# equivalent: select(starwars2, -birth_year, -c(films:starships))
starwars3
head(select(starwars, ends_with("color")))
head(select(starwars, starts_with("s")))
- Notice that
select()may not work properly if theMASSpackage is attached. If that happens, detachMASSin the Packages pane or rundetach("package:MASS", unload = TRUE).
Rename columns with rename()
- Renames columns using
new_name = old_name.
rename(.data, ...)
.data: a data frame or tibble.
...: use new_name = old_name to rename selected variables.
starwars4 = rename(
starwars3,
haircolor = hair_color,
skincolor = skin_color,
eyecolor = eye_color
)
starwars4
Modify/Add columns with mutate()
- Modifies an existing column.
- Creates a new column if it does not exist.
mutate(.data, ...)
.data: a data frame or tibble.
...: name-value pairs defining the output columns.
starwars5 = mutate(
starwars4,
height = height / 100, # convert cm to meters
BMI = mass / height^2,
dummy = 1 # a scalar is recycled to all rows
)
starwars5 = select(starwars5, BMI, dummy, everything())
starwars5
The pipe operator %>%
- Notice that all previous
dplyrfunctions take the dataset as their first argument. - The pipe operator
%>%passes the object on the left-hand side into the first argument of the function on the right-hand side.
filter(starwars, species == "Droid") # without the pipe
starwars %>% filter(species == "Droid") # with the pipe
- When we use the pipe operator, the dataset should not be written again as the first argument because
%>%passes it automatically. - Also note that, from
filter()throughmutate(), we have been accumulating transformations in new data frames. The last object,starwars5, is the result of applying the full sequence of changes to the originalstarwarsdataset.
starwars1 = filter(starwars, species == "Human", height >= 100)
starwars2 = arrange(starwars1, height, desc(mass))
starwars3 = select(starwars2, name:eye_color, sex:species)
starwars4 = rename(
starwars3,
haircolor = hair_color,
skincolor = skin_color,
eyecolor = eye_color
)
starwars5 = mutate(
starwars4,
height = height / 100,
BMI = mass / height^2,
dummy = 1
)
starwars5 = select(starwars5, BMI, dummy, everything())
starwars5
- By chaining the same commands with
%>%, we can write the full workflow as a single pipeline:
starwars_pipe = starwars %>%
filter(species == "Human", height >= 100) %>%
arrange(height, desc(mass)) %>%
select(name:eye_color, sex:species) %>%
rename(
haircolor = hair_color,
skincolor = skin_color,
eyecolor = eye_color
) %>%
mutate(
height = height / 100,
BMI = mass / height^2,
dummy = 1
) %>%
select(BMI, dummy, everything())
starwars_pipe
all(starwars_pipe == starwars5, na.rm = TRUE)
Summarize with summarise()
- We can use
summarise()to compute statistics for one or more variables:
starwars %>% summarise(
n_obs = n(),
mean_height = mean(height, na.rm = TRUE),
mean_mass = mean(mass, na.rm = TRUE)
)
- In the example above, we simply computed the sample size and the mean height and mass for the full sample.
Group data with group_by()
- Unlike the other
dplyrfunctions shown so far,group_by()does not change the contents of the data frame. It only turns the data into a grouped dataset according to the categories of one or more variables.
grouped_sw = starwars %>% group_by(sex)
class(grouped_sw)
head(starwars)
head(grouped_sw)
group_by()is useful whenever the next operation depends on multiple rows. As an illustration, let us create a column with the mean value ofmassacross all observations:
starwars %>%
mutate(mean_mass = mean(mass, na.rm = TRUE)) %>%
select(mean_mass, sex, everything()) %>%
head(10)
- Notice that all values of
mean_massare identical. We now group bysexbefore computing the mean:
starwars %>%
group_by(sex) %>%
mutate(mean_mass = mean(mass, na.rm = TRUE)) %>%
ungroup() %>%
select(mean_mass, sex, everything()) %>%
head(10)
- This is useful in many economics applications in which we work with group-level variables, for example household-level statistics attached to individual observations.
Avoid mistakes: whenever you use
group_by(), remember to runungroup()after the grouped operation if the next commands should not remain grouped.
Grouped summaries with group_by() and summarise()
summarise()becomes especially useful when combined withgroup_by(), because it lets us compute statistics at the group level:
summary_sw = starwars %>%
group_by(sex) %>%
summarise(
n_obs = n(),
mean_height = mean(height, na.rm = TRUE),
mean_mass = mean(mass, na.rm = TRUE)
)
summary_sw
class(summary_sw)
- After
summarise(), the resulting data frame is no longer grouped, soungroup()is not needed in this case. - We can also group by more than one variable:
starwars %>%
group_by(sex, hair_color) %>%
summarise(
n_obs = n(),
mean_height = mean(height, na.rm = TRUE),
mean_mass = mean(mass, na.rm = TRUE)
)
- To group continuous variables, we first need to define intervals with
cut():
cut(x, ...)
x: a numeric vector that will be turned into a factor by cutting it into intervals.
breaks: either a vector of cut points or a single integer giving the number of intervals.
# breaks supplied as an integer = desired number of groups
starwars %>%
group_by(cut(birth_year, breaks = 5)) %>%
summarise(
n_obs = n(),
mean_height = mean(height, na.rm = TRUE)
)
# breaks supplied as a vector = explicit cut points
starwars %>%
group_by(birth_year = cut(birth_year, breaks = c(0, 40, 90, 200, 900))) %>%
summarise(
n_obs = n(),
mean_height = mean(height, na.rm = TRUE)
)
- Notice that we wrote
birth_year = cut(birth_year, ...)so the grouped column keeps the namebirth_year.
Join datasets with _join_ functions
- Earlier we saw that
merge()combines datasets through key variables. - The
dplyrpackage provides a family of_join_functions that accomplish the same task with a cleaner interface:
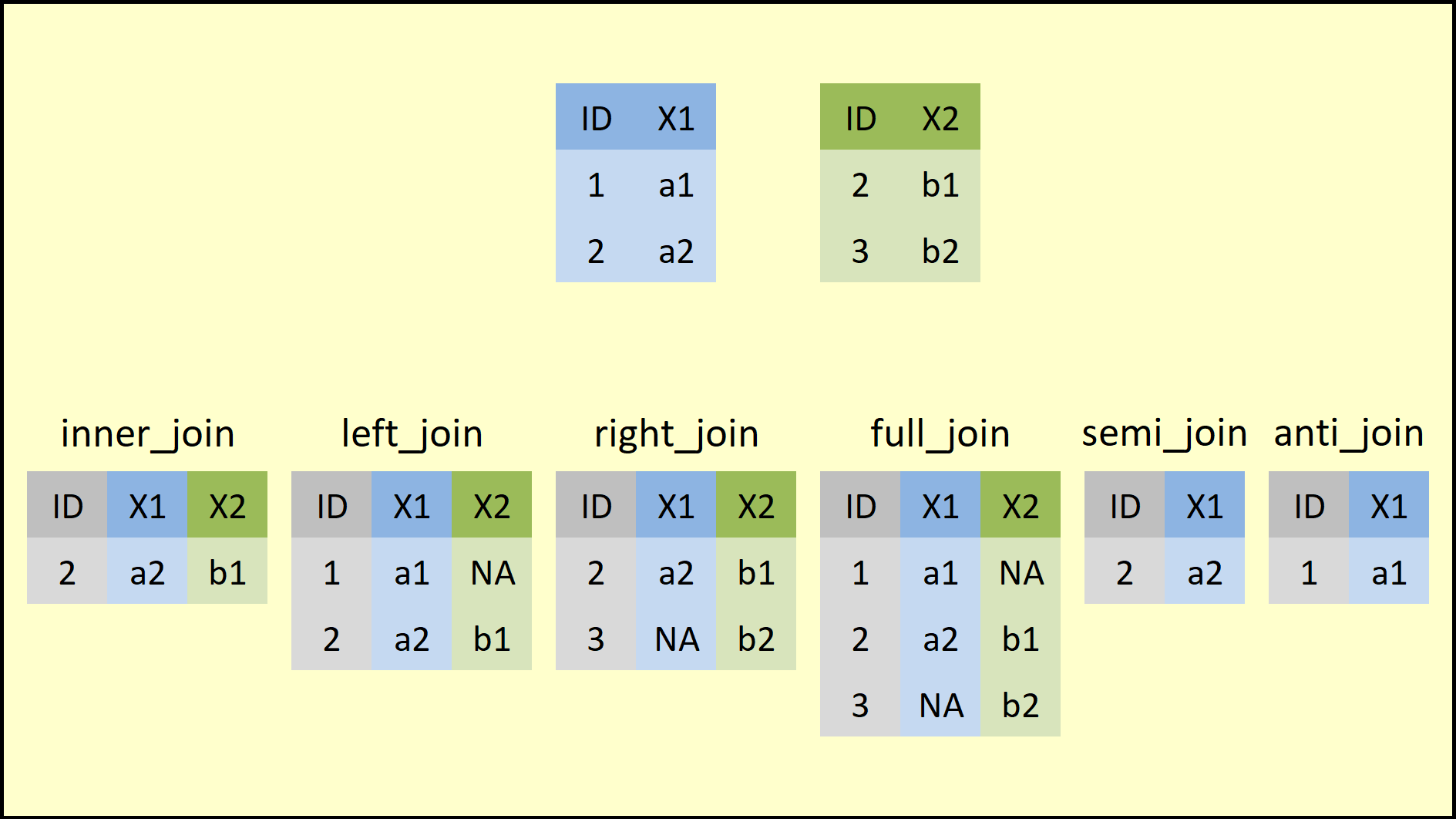
The common ingredients are:
x: dataset 1y: dataset 2by: vector of key variablessuffix: suffixes used when non-key variables share the same name
As an example, we use subsets of the
starwarsdataset:
bd1 = starwars[1:6, c(1, 3, 11)]
bd2 = starwars[c(2, 4, 7:10), c(1:2, 6)]
bd1
bd2
There are 12 unique characters across the two datasets, but only
"C-3PO"and"Darth Vader"appear in both.inner_join(): keeps only IDs present in both datasets
inner_join(bd1, bd2, by = "name")
full_join(): keeps all IDs, even if they appear in only one dataset
full_join(bd1, bd2, by = "name")
left_join(): keeps the IDs present in dataset 1, supplied asx
left_join(bd1, bd2, by = "name")
right_join(): keeps the IDs present in dataset 2, supplied asy
right_join(bd1, bd2, by = "name")