The dplyr package
- Vignette - Introduction to dplyr
- The
dplyrpackage makes data manipulation easier through simple and computationally efficient functions. - These functions can be grouped into three categories:
- Columns:
select(): selects or drops data-frame columnsrename(): changes column namesmutate(): creates or modifies column values
- Rows:
filter(): selects rows according to column valuesarrange(): reorders the rows
- Groups of rows:
summarise(): collapses each group into a single row
- Columns:
- In this subsection, we continue using the Star Wars dataset (
starwars) introduced in the previous subsection. - You will notice that, when these functions are applied, the data frame is turned into a tibble, which is a more efficient format for tabular data but works very similarly to a standard data frame.
library("dplyr") # Load package
##
## Attaching package: 'dplyr'
## The following objects are masked from 'package:stats':
##
## filter, lag
## The following objects are masked from 'package:base':
##
## intersect, setdiff, setequal, union
head(starwars) # inspect the first rows of the dataset included in the package
## # A tibble: 6 × 14
## name height mass hair_color skin_color eye_color birth_year sex gender
## <chr> <int> <dbl> <chr> <chr> <chr> <dbl> <chr> <chr>
## 1 Luke Sky… 172 77 blond fair blue 19 male mascu…
## 2 C-3PO 167 75 <NA> gold yellow 112 none mascu…
## 3 R2-D2 96 32 <NA> white, bl… red 33 none mascu…
## 4 Darth Va… 202 136 none white yellow 41.9 male mascu…
## 5 Leia Org… 150 49 brown light brown 19 fema… femin…
## 6 Owen Lars 178 120 brown, gr… light blue 52 male mascu…
## # ℹ 5 more variables: homeworld <chr>, species <chr>, films <list>,
## # vehicles <list>, starships <list>
Filter rows with filter()
- Selects a subset of rows from a data frame.
filter(.data, ...)
.data: A data frame, data frame extension (e.g. a tibble), or a lazy data frame (e.g. from dbplyr or dtplyr).
... : <data-masking> Expressions that return a logical value, and are defined in terms of the variables in .data. If multiple expressions are included, they are combined with the & operator. Only rows for which all conditions evaluate to TRUE are kept.
starwars1 = filter(starwars, species == "Human", height >= 100)
starwars1
## # A tibble: 31 × 14
## name height mass hair_color skin_color eye_color birth_year sex gender
## <chr> <int> <dbl> <chr> <chr> <chr> <dbl> <chr> <chr>
## 1 Luke Sk… 172 77 blond fair blue 19 male mascu…
## 2 Darth V… 202 136 none white yellow 41.9 male mascu…
## 3 Leia Or… 150 49 brown light brown 19 fema… femin…
## 4 Owen La… 178 120 brown, gr… light blue 52 male mascu…
## 5 Beru Wh… 165 75 brown light blue 47 fema… femin…
## 6 Biggs D… 183 84 black light brown 24 male mascu…
## 7 Obi-Wan… 182 77 auburn, w… fair blue-gray 57 male mascu…
## 8 Anakin … 188 84 blond fair blue 41.9 male mascu…
## 9 Wilhuff… 180 NA auburn, g… fair blue 64 male mascu…
## 10 Han Solo 180 80 brown fair brown 29 male mascu…
## # ℹ 21 more rows
## # ℹ 5 more variables: homeworld <chr>, species <chr>, films <list>,
## # vehicles <list>, starships <list>
# Equivalente a:
starwars[starwars$species == "Human" & starwars$height >= 100, ]
## # A tibble: 39 × 14
## name height mass hair_color skin_color eye_color birth_year sex gender
## <chr> <int> <dbl> <chr> <chr> <chr> <dbl> <chr> <chr>
## 1 Luke Sk… 172 77 blond fair blue 19 male mascu…
## 2 Darth V… 202 136 none white yellow 41.9 male mascu…
## 3 Leia Or… 150 49 brown light brown 19 fema… femin…
## 4 Owen La… 178 120 brown, gr… light blue 52 male mascu…
## 5 Beru Wh… 165 75 brown light blue 47 fema… femin…
## 6 Biggs D… 183 84 black light brown 24 male mascu…
## 7 Obi-Wan… 182 77 auburn, w… fair blue-gray 57 male mascu…
## 8 Anakin … 188 84 blond fair blue 41.9 male mascu…
## 9 Wilhuff… 180 NA auburn, g… fair blue 64 male mascu…
## 10 Han Solo 180 80 brown fair brown 29 male mascu…
## # ℹ 29 more rows
## # ℹ 5 more variables: homeworld <chr>, species <chr>, films <list>,
## # vehicles <list>, starships <list>
Reorder rows with arrange()
- Reorders the rows according to one or more column names.
arrange(.data, ..., .by_group = FALSE)
.data: A data frame, data frame extension (e.g. a tibble), or a lazy data frame (e.g. from dbplyr or dtplyr).
... : <data-masking> Variables, or functions of variables. Use desc() to sort a variable in descending order.
- If more than one variable is provided, the rows are first sorted by the first variable and ties are broken using the second variable.
- To sort in descending order, use the
desc()function.
starwars2 = arrange(starwars1, height, desc(mass))
starwars2
## # A tibble: 31 × 14
## name height mass hair_color skin_color eye_color birth_year sex gender
## <chr> <int> <dbl> <chr> <chr> <chr> <dbl> <chr> <chr>
## 1 Leia Or… 150 49 brown light brown 19 fema… femin…
## 2 Mon Mot… 150 NA auburn fair blue 48 fema… femin…
## 3 Cordé 157 NA brown light brown NA fema… femin…
## 4 Shmi Sk… 163 NA black fair brown 72 fema… femin…
## 5 Beru Wh… 165 75 brown light blue 47 fema… femin…
## 6 Padmé A… 165 45 brown light brown 46 fema… femin…
## 7 Dormé 165 NA brown light brown NA fema… femin…
## 8 Jocasta… 167 NA white fair blue NA fema… femin…
## 9 Wedge A… 170 77 brown fair hazel 21 male mascu…
## 10 Palpati… 170 75 grey pale yellow 82 male mascu…
## # ℹ 21 more rows
## # ℹ 5 more variables: homeworld <chr>, species <chr>, films <list>,
## # vehicles <list>, starships <list>
Select columns with select()
- Selects the columns of interest.
select(.data, ...)
... : variables in a data frame
: for selecting a range of consecutive variables.
! for taking the complement of a set of variables.
c() for combining selections.
- To select columns, simply list the desired column names.
- We can also select a range of variables with
var1:var2. - If we want to drop only a few columns, we can list their names preceded by a minus sign (
-var).
starwars3 = select(starwars2, name:eye_color, sex:species)
# equivalente: select(starwars2, -birth_year, -c(films:starships))
starwars3
## # A tibble: 31 × 10
## name height mass hair_color skin_color eye_color sex gender homeworld
## <chr> <int> <dbl> <chr> <chr> <chr> <chr> <chr> <chr>
## 1 Leia Org… 150 49 brown light brown fema… femin… Alderaan
## 2 Mon Moth… 150 NA auburn fair blue fema… femin… Chandrila
## 3 Cordé 157 NA brown light brown fema… femin… Naboo
## 4 Shmi Sky… 163 NA black fair brown fema… femin… Tatooine
## 5 Beru Whi… 165 75 brown light blue fema… femin… Tatooine
## 6 Padmé Am… 165 45 brown light brown fema… femin… Naboo
## 7 Dormé 165 NA brown light brown fema… femin… Naboo
## 8 Jocasta … 167 NA white fair blue fema… femin… Coruscant
## 9 Wedge An… 170 77 brown fair hazel male mascu… Corellia
## 10 Palpatine 170 75 grey pale yellow male mascu… Naboo
## # ℹ 21 more rows
## # ℹ 1 more variable: species <chr>
- Notice that
select()may not work properly if theMASSpackage is attached. If that happens, detachMASSin the lower-right Packages pane or rundetach("package:MASS", unload = TRUE). - Another way to select columns is to combine
select()with helpers such asstarts_with()andends_with(), which select columns that begin or end with a given text pattern.
head( select(starwars, ends_with("color")) ) # colunas que terminam com color
## # A tibble: 6 × 3
## hair_color skin_color eye_color
## <chr> <chr> <chr>
## 1 blond fair blue
## 2 <NA> gold yellow
## 3 <NA> white, blue red
## 4 none white yellow
## 5 brown light brown
## 6 brown, grey light blue
head( select(starwars, starts_with("s")) ) # colunas que iniciam com a letra "s"
## # A tibble: 6 × 4
## skin_color sex species starships
## <chr> <chr> <chr> <list>
## 1 fair male Human <chr [2]>
## 2 gold none Droid <chr [0]>
## 3 white, blue none Droid <chr [0]>
## 4 white male Human <chr [1]>
## 5 light female Human <chr [0]>
## 6 light male Human <chr [0]>
Rename columns with rename()
- Renames columns using
new_name = old_name.
rename(.data, ...)
.data: A data frame, data frame extension (e.g. a tibble), or a lazy data frame (e.g. from dbplyr or dtplyr).
... : For rename(): <tidy-select> Use new_name = old_name to rename selected variables.
starwars4 = rename(starwars3,
haircolor = hair_color,
skincolor = skin_color,
eyecolor = eye_color)
starwars4
## # A tibble: 31 × 10
## name height mass haircolor skincolor eyecolor sex gender homeworld
## <chr> <int> <dbl> <chr> <chr> <chr> <chr> <chr> <chr>
## 1 Leia Organa 150 49 brown light brown fema… femin… Alderaan
## 2 Mon Mothma 150 NA auburn fair blue fema… femin… Chandrila
## 3 Cordé 157 NA brown light brown fema… femin… Naboo
## 4 Shmi Skywal… 163 NA black fair brown fema… femin… Tatooine
## 5 Beru Whites… 165 75 brown light blue fema… femin… Tatooine
## 6 Padmé Amida… 165 45 brown light brown fema… femin… Naboo
## 7 Dormé 165 NA brown light brown fema… femin… Naboo
## 8 Jocasta Nu 167 NA white fair blue fema… femin… Coruscant
## 9 Wedge Antil… 170 77 brown fair hazel male mascu… Corellia
## 10 Palpatine 170 75 grey pale yellow male mascu… Naboo
## # ℹ 21 more rows
## # ℹ 1 more variable: species <chr>
Modify/Add columns with mutate()
- Modifies an existing column
- Creates a new column if it does not exist
mutate(.data, ...)
.data: A data frame, data frame extension (e.g. a tibble), or a lazy data frame (e.g. from dbplyr or dtplyr).
... : <data-masking> Name-value pairs. The name gives the name of the column in the output. The value can be:
- A vector of length 1, which will be recycled to the correct length.
- A vector the same length as the current group (or the whole data frame if ungrouped).
- NULL, to remove the column.
starwars5 = mutate(starwars4,
height = height/100, # convert cm to meters
BMI = mass / height^2,
dummy = 1 # if it is not a vector, the same value is recycled
)
starwars5 = select(starwars5, BMI, dummy, everything()) # make reading easier
starwars5
## # A tibble: 31 × 12
## BMI dummy name height mass haircolor skincolor eyecolor sex gender
## <dbl> <dbl> <chr> <dbl> <dbl> <chr> <chr> <chr> <chr> <chr>
## 1 21.8 1 Leia Orga… 1.5 49 brown light brown fema… femin…
## 2 NA 1 Mon Mothma 1.5 NA auburn fair blue fema… femin…
## 3 NA 1 Cordé 1.57 NA brown light brown fema… femin…
## 4 NA 1 Shmi Skyw… 1.63 NA black fair brown fema… femin…
## 5 27.5 1 Beru Whit… 1.65 75 brown light blue fema… femin…
## 6 16.5 1 Padmé Ami… 1.65 45 brown light brown fema… femin…
## 7 NA 1 Dormé 1.65 NA brown light brown fema… femin…
## 8 NA 1 Jocasta Nu 1.67 NA white fair blue fema… femin…
## 9 26.6 1 Wedge Ant… 1.7 77 brown fair hazel male mascu…
## 10 26.0 1 Palpatine 1.7 75 grey pale yellow male mascu…
## # ℹ 21 more rows
## # ℹ 2 more variables: homeworld <chr>, species <chr>
The pipe operator %>%
- Notice that all previous
dplyrfunctions take the dataset (.data) as their first argument, and that is not accidental. - The pipe operator
%>%passes the data frame written on the left-hand side into the first argument of the next function on the right-hand side.
filter(starwars, species=="Droid") # without the pipe operator
## # A tibble: 6 × 14
## name height mass hair_color skin_color eye_color birth_year sex gender
## <chr> <int> <dbl> <chr> <chr> <chr> <dbl> <chr> <chr>
## 1 C-3PO 167 75 <NA> gold yellow 112 none masculi…
## 2 R2-D2 96 32 <NA> white, blue red 33 none masculi…
## 3 R5-D4 97 32 <NA> white, red red NA none masculi…
## 4 IG-88 200 140 none metal red 15 none masculi…
## 5 R4-P17 96 NA none silver, red red, blue NA none feminine
## 6 BB8 NA NA none none black NA none masculi…
## # ℹ 5 more variables: homeworld <chr>, species <chr>, films <list>,
## # vehicles <list>, starships <list>
starwars %>% filter(species=="Droid") # with the pipe operator
## # A tibble: 6 × 14
## name height mass hair_color skin_color eye_color birth_year sex gender
## <chr> <int> <dbl> <chr> <chr> <chr> <dbl> <chr> <chr>
## 1 C-3PO 167 75 <NA> gold yellow 112 none masculi…
## 2 R2-D2 96 32 <NA> white, blue red 33 none masculi…
## 3 R5-D4 97 32 <NA> white, red red NA none masculi…
## 4 IG-88 200 140 none metal red 15 none masculi…
## 5 R4-P17 96 NA none silver, red red, blue NA none feminine
## 6 BB8 NA NA none none black NA none masculi…
## # ℹ 5 more variables: homeworld <chr>, species <chr>, films <list>,
## # vehicles <list>, starships <list>
- Notice that, when we use the pipe operator, the first argument containing the dataset should not be filled in, because it is already passed automatically by
%>%. - Also note that, from the subsection on
filter()throughmutate(), we have been accumulating the transformations in new data frames. In other words, the last object,starwars5, is the originalstarwarsdataset after the sequence of changes produced byfilter(),arrange(),select(),rename(), andmutate().
starwars1 = filter(starwars, species == "Human", height >= 100)
starwars2 = arrange(starwars1, height, desc(mass))
starwars3 = select(starwars2, name:eye_color, sex:species)
starwars4 = rename(starwars3,
haircolor = hair_color,
skincolor = skin_color,
eyecolor = eye_color)
starwars5 = mutate(starwars4,
height = height/100,
BMI = mass / height^2,
dummy = 1
)
starwars5 = select(starwars5, BMI, dummy, everything())
starwars5
## # A tibble: 31 × 12
## BMI dummy name height mass haircolor skincolor eyecolor sex gender
## <dbl> <dbl> <chr> <dbl> <dbl> <chr> <chr> <chr> <chr> <chr>
## 1 21.8 1 Leia Orga… 1.5 49 brown light brown fema… femin…
## 2 NA 1 Mon Mothma 1.5 NA auburn fair blue fema… femin…
## 3 NA 1 Cordé 1.57 NA brown light brown fema… femin…
## 4 NA 1 Shmi Skyw… 1.63 NA black fair brown fema… femin…
## 5 27.5 1 Beru Whit… 1.65 75 brown light blue fema… femin…
## 6 16.5 1 Padmé Ami… 1.65 45 brown light brown fema… femin…
## 7 NA 1 Dormé 1.65 NA brown light brown fema… femin…
## 8 NA 1 Jocasta Nu 1.67 NA white fair blue fema… femin…
## 9 26.6 1 Wedge Ant… 1.7 77 brown fair hazel male mascu…
## 10 26.0 1 Palpatine 1.7 75 grey pale yellow male mascu…
## # ℹ 21 more rows
## # ℹ 2 more variables: homeworld <chr>, species <chr>
- By using the pipe operator
%>%repeatedly, we can take the output from one function and feed it directly into the next one. We can therefore rewrite the code above as a single pipeline and obtain the same data frame asstarwars5.
starwars_pipe = starwars %>%
filter(species == "Human", height >= 100) %>%
arrange(height, desc(mass)) %>%
select(name:eye_color, sex:species) %>%
rename(haircolor = hair_color,
skincolor = skin_color,
eyecolor = eye_color) %>%
mutate(height = height/100,
BMI = mass / height^2,
dummy = 1
) %>%
select(BMI, dummy, everything())
starwars_pipe
## # A tibble: 31 × 12
## BMI dummy name height mass haircolor skincolor eyecolor sex gender
## <dbl> <dbl> <chr> <dbl> <dbl> <chr> <chr> <chr> <chr> <chr>
## 1 21.8 1 Leia Orga… 1.5 49 brown light brown fema… femin…
## 2 NA 1 Mon Mothma 1.5 NA auburn fair blue fema… femin…
## 3 NA 1 Cordé 1.57 NA brown light brown fema… femin…
## 4 NA 1 Shmi Skyw… 1.63 NA black fair brown fema… femin…
## 5 27.5 1 Beru Whit… 1.65 75 brown light blue fema… femin…
## 6 16.5 1 Padmé Ami… 1.65 45 brown light brown fema… femin…
## 7 NA 1 Dormé 1.65 NA brown light brown fema… femin…
## 8 NA 1 Jocasta Nu 1.67 NA white fair blue fema… femin…
## 9 26.6 1 Wedge Ant… 1.7 77 brown fair hazel male mascu…
## 10 26.0 1 Palpatine 1.7 75 grey pale yellow male mascu…
## # ℹ 21 more rows
## # ℹ 2 more variables: homeworld <chr>, species <chr>
all(starwars_pipe == starwars5, na.rm=TRUE) # check whether all elements are equal
## [1] TRUE
Resuma com summarise()
- We can use
summarise()to compute statistics for one or more variables:
starwars %>% summarise(
n_obs = n(),
mean_height = mean(height, na.rm=TRUE),
mean_mass = mean(mass, na.rm=TRUE)
)
## # A tibble: 1 × 3
## n_obs mean_height mean_mass
## <int> <dbl> <dbl>
## 1 87 174. 97.3
- In the example above, we simply obtained the sample size and the mean height and mass for the full
starwarssample, which is not especially informative.
Agrupe com group_by()
- Unlike the other
dplyrfunctions shown so far, the output ofgroup_by()does not change the content of the data frame. It only turns the data into a grouped dataset according to the categories of a given variable.
grouped_sw = starwars %>% group_by(sex)
class(grouped_sw)
## [1] "grouped_df" "tbl_df" "tbl" "data.frame"
head(starwars)
## # A tibble: 6 × 14
## name height mass hair_color skin_color eye_color birth_year sex gender
## <chr> <int> <dbl> <chr> <chr> <chr> <dbl> <chr> <chr>
## 1 Luke Sky… 172 77 blond fair blue 19 male mascu…
## 2 C-3PO 167 75 <NA> gold yellow 112 none mascu…
## 3 R2-D2 96 32 <NA> white, bl… red 33 none mascu…
## 4 Darth Va… 202 136 none white yellow 41.9 male mascu…
## 5 Leia Org… 150 49 brown light brown 19 fema… femin…
## 6 Owen Lars 178 120 brown, gr… light blue 52 male mascu…
## # ℹ 5 more variables: homeworld <chr>, species <chr>, films <list>,
## # vehicles <list>, starships <list>
head(grouped_sw) # grouped by sex
## # A tibble: 6 × 14
## # Groups: sex [3]
## name height mass hair_color skin_color eye_color birth_year sex gender
## <chr> <int> <dbl> <chr> <chr> <chr> <dbl> <chr> <chr>
## 1 Luke Sky… 172 77 blond fair blue 19 male mascu…
## 2 C-3PO 167 75 <NA> gold yellow 112 none mascu…
## 3 R2-D2 96 32 <NA> white, bl… red 33 none mascu…
## 4 Darth Va… 202 136 none white yellow 41.9 male mascu…
## 5 Leia Org… 150 49 brown light brown 19 fema… femin…
## 6 Owen Lars 178 120 brown, gr… light blue 52 male mascu…
## # ℹ 5 more variables: homeworld <chr>, species <chr>, films <list>,
## # vehicles <list>, starships <list>
group_by()prepares the data frame for operations that depend on multiple rows. As an example, let us create a column containing the mean ofmassacross all observations.
starwars %>%
mutate(mean_mass = mean(mass, na.rm=TRUE)) %>%
select(mean_mass, sex, everything()) %>%
head(10)
## # A tibble: 10 × 15
## mean_mass sex name height mass hair_color skin_color eye_color birth_year
## <dbl> <chr> <chr> <int> <dbl> <chr> <chr> <chr> <dbl>
## 1 97.3 male Luke… 172 77 blond fair blue 19
## 2 97.3 none C-3PO 167 75 <NA> gold yellow 112
## 3 97.3 none R2-D2 96 32 <NA> white, bl… red 33
## 4 97.3 male Dart… 202 136 none white yellow 41.9
## 5 97.3 fema… Leia… 150 49 brown light brown 19
## 6 97.3 male Owen… 178 120 brown, gr… light blue 52
## 7 97.3 fema… Beru… 165 75 brown light blue 47
## 8 97.3 none R5-D4 97 32 <NA> white, red red NA
## 9 97.3 male Bigg… 183 84 black light brown 24
## 10 97.3 male Obi-… 182 77 auburn, w… fair blue-gray 57
## # ℹ 6 more variables: gender <chr>, homeworld <chr>, species <chr>,
## # films <list>, vehicles <list>, starships <list>
- Notice that all values of
mean_massare identical. We now group bysexbefore computing the mean:
starwars %>%
group_by(sex) %>%
mutate(mean_mass = mean(mass, na.rm=TRUE)) %>%
ungroup() %>% # always remember to ungroup afterward
select(mean_mass, sex, everything()) %>%
head(10)
## # A tibble: 10 × 15
## mean_mass sex name height mass hair_color skin_color eye_color birth_year
## <dbl> <chr> <chr> <int> <dbl> <chr> <chr> <chr> <dbl>
## 1 81.0 male Luke… 172 77 blond fair blue 19
## 2 69.8 none C-3PO 167 75 <NA> gold yellow 112
## 3 69.8 none R2-D2 96 32 <NA> white, bl… red 33
## 4 81.0 male Dart… 202 136 none white yellow 41.9
## 5 54.7 fema… Leia… 150 49 brown light brown 19
## 6 81.0 male Owen… 178 120 brown, gr… light blue 52
## 7 54.7 fema… Beru… 165 75 brown light blue 47
## 8 69.8 none R5-D4 97 32 <NA> white, red red NA
## 9 81.0 male Bigg… 183 84 black light brown 24
## 10 81.0 male Obi-… 182 77 auburn, w… fair blue-gray 57
## # ℹ 6 more variables: gender <chr>, homeworld <chr>, species <chr>,
## # films <list>, vehicles <list>, starships <list>
- Notice that the
mean_masscolumn now takes different values depending on the sex of the observation. - This is useful in many economic applications where we work with group-level variables, for example at the household level, attached to individual observations such as household members.
Avoid potential mistakes: Whenever you use
group_by(), remember to ungroup the data frame withungroup()after carrying out the desired operations.
Resuma em grupos com group_by() e summarise()
summarise()is especially useful when combined withgroup_by(), because it allows us to compute group-level statistics:
summary_sw = starwars %>% group_by(sex) %>%
summarise(
n_obs = n(),
mean_height = mean(height, na.rm = TRUE),
mean_mass = mean(mass, na.rm = TRUE)
)
summary_sw
## # A tibble: 5 × 4
## sex n_obs mean_height mean_mass
## <chr> <int> <dbl> <dbl>
## 1 female 16 169. 54.7
## 2 hermaphroditic 1 175 1358
## 3 male 60 179. 81.0
## 4 none 6 131. 69.8
## 5 <NA> 4 181. 48
class(summary_sw) # once we summarise, the result is no longer grouped
## [1] "tbl_df" "tbl" "data.frame"
- Notice that, after using
summarise(), the resulting data frame is no longer grouped, soungroup()is not necessary in this case. - We can also group by more than one variable:
starwars %>% group_by(sex, hair_color) %>%
summarise(
n_obs = n(),
mean_height = mean(height, na.rm = TRUE),
mean_mass = mean(mass, na.rm = TRUE)
)
## `summarise()` has grouped output by 'sex'. You can override using the `.groups`
## argument.
## # A tibble: 23 × 5
## # Groups: sex [5]
## sex hair_color n_obs mean_height mean_mass
## <chr> <chr> <int> <dbl> <dbl>
## 1 female auburn 1 150 NaN
## 2 female black 3 166. 53.1
## 3 female blonde 1 168 55
## 4 female brown 6 160. 56.3
## 5 female none 4 188. 54
## 6 female white 1 167 NaN
## 7 hermaphroditic <NA> 1 175 1358
## 8 male auburn, grey 1 180 NaN
## 9 male auburn, white 1 182 77
## 10 male black 9 176. 81.0
## # ℹ 13 more rows
- To group continuous variables, we need to define intervals with the
cut()function.
cut(x, ...)
x: a numeric vector which is to be converted to a factor by cutting.
breaks: either a numeric vector of two or more unique cut points or a single number (greater than or equal to 2) giving the number of intervals into which x is to be cut.
# breaks supplied as an integer = desired number of groups
starwars %>% group_by(cut(birth_year, breaks=5)) %>%
summarise(
n_obs = n(),
mean_height = mean(height, na.rm = TRUE)
)
## # A tibble: 5 × 3
## `cut(birth_year, breaks = 5)` n_obs mean_height
## <fct> <int> <dbl>
## 1 (7.11,186] 40 175.
## 2 (186,363] 1 228
## 3 (541,718] 1 175
## 4 (718,897] 1 66
## 5 <NA> 44 176.
# breaks supplied as a vector = group cut points
starwars %>% group_by(birth_year=cut(birth_year,
breaks=c(0, 40, 90, 200, 900))) %>%
summarise(
n_obs = n(),
mean_height = mean(height, na.rm = TRUE)
)
## # A tibble: 5 × 3
## birth_year n_obs mean_height
## <fct> <int> <dbl>
## 1 (0,40] 13 164.
## 2 (40,90] 22 179.
## 3 (90,200] 6 192.
## 4 (200,900] 2 120.
## 5 <NA> 44 176.
- Notice that we wrote
birth_year = cut(birth_year, ...)so the column keeps the namebirth_year; otherwise it would be namedcut(birth_year, ...).
Join datasets with join functions
- We saw earlier that
cbind()can be used to combine one data frame with another data frame, or a vector, as long as they have the same number of rows. - To append rows, provided the columns share compatible classes, we can use
rbind(). - To combine datasets using key variables, we use
merge(). - The
dplyrpackage provides a family of join functions that accomplish the same task asmerge(). Instead of changing an argument value, however, we choose the specific join function that matches the merge we want, as summarized in the figure below:
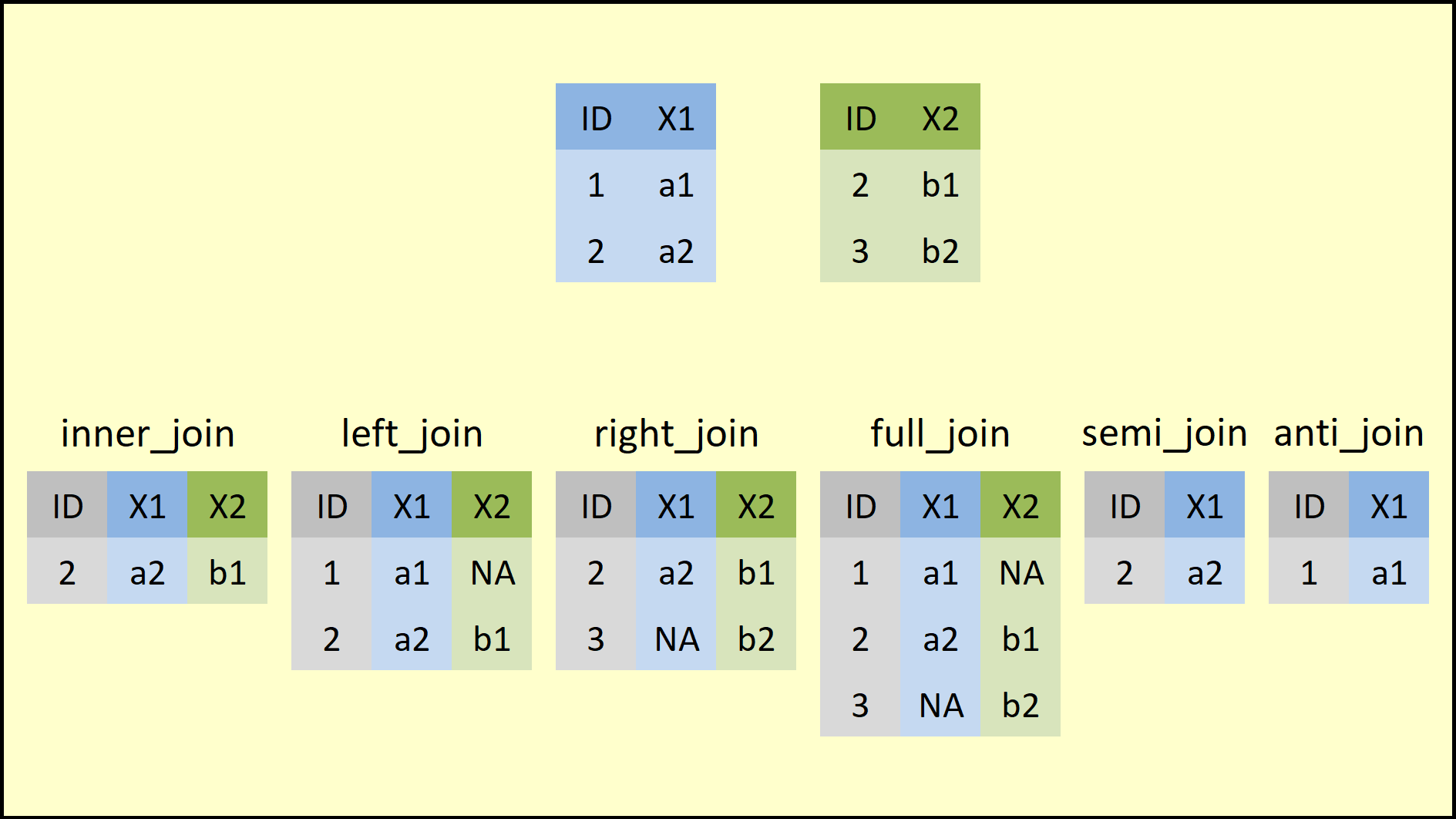
- All these functions share the same basic syntax:
x: dataset 1y: dataset 2by: vector of key variablessuffix: vector with two suffixes used for columns that share the same name
- As an example, we use subsets of the
starwarsdataset:
bd1 = starwars[1:6, c(1, 3, 11)]
bd2 = starwars[c(2, 4, 7:10), c(1:2, 6)]
bd1
## # A tibble: 6 × 3
## name mass species
## <chr> <dbl> <chr>
## 1 Luke Skywalker 77 Human
## 2 C-3PO 75 Droid
## 3 R2-D2 32 Droid
## 4 Darth Vader 136 Human
## 5 Leia Organa 49 Human
## 6 Owen Lars 120 Human
bd2
## # A tibble: 6 × 3
## name height eye_color
## <chr> <int> <chr>
## 1 C-3PO 167 yellow
## 2 Darth Vader 202 yellow
## 3 Beru Whitesun lars 165 blue
## 4 R5-D4 97 red
## 5 Biggs Darklighter 183 brown
## 6 Obi-Wan Kenobi 182 blue-gray
- Notice that there are 12 unique characters across the two datasets, but only
"C-3PO"and"Darth Vader"appear in both datasets. inner_join(): keeps only IDs present in both datasets
inner_join(bd1, bd2, by="name")
## # A tibble: 2 × 5
## name mass species height eye_color
## <chr> <dbl> <chr> <int> <chr>
## 1 C-3PO 75 Droid 167 yellow
## 2 Darth Vader 136 Human 202 yellow
full_join(): keeps all IDs, even if they appear in only one of the two datasets
full_join(bd1, bd2, by="name")
## # A tibble: 10 × 5
## name mass species height eye_color
## <chr> <dbl> <chr> <int> <chr>
## 1 Luke Skywalker 77 Human NA <NA>
## 2 C-3PO 75 Droid 167 yellow
## 3 R2-D2 32 Droid NA <NA>
## 4 Darth Vader 136 Human 202 yellow
## 5 Leia Organa 49 Human NA <NA>
## 6 Owen Lars 120 Human NA <NA>
## 7 Beru Whitesun lars NA <NA> 165 blue
## 8 R5-D4 NA <NA> 97 red
## 9 Biggs Darklighter NA <NA> 183 brown
## 10 Obi-Wan Kenobi NA <NA> 182 blue-gray
left_join(): keeps the IDs present in dataset 1 (supplied asx)
left_join(bd1, bd2, by="name")
## # A tibble: 6 × 5
## name mass species height eye_color
## <chr> <dbl> <chr> <int> <chr>
## 1 Luke Skywalker 77 Human NA <NA>
## 2 C-3PO 75 Droid 167 yellow
## 3 R2-D2 32 Droid NA <NA>
## 4 Darth Vader 136 Human 202 yellow
## 5 Leia Organa 49 Human NA <NA>
## 6 Owen Lars 120 Human NA <NA>
right_join(): keeps the IDs present in dataset 2 (supplied asy)
right_join(bd1, bd2, by="name")
## # A tibble: 6 × 5
## name mass species height eye_color
## <chr> <dbl> <chr> <int> <chr>
## 1 C-3PO 75 Droid 167 yellow
## 2 Darth Vader 136 Human 202 yellow
## 3 Beru Whitesun lars NA <NA> 165 blue
## 4 R5-D4 NA <NA> 97 red
## 5 Biggs Darklighter NA <NA> 183 brown
## 6 Obi-Wan Kenobi NA <NA> 182 blue-gray
- We can also use more than one key variable to match IDs across datasets. First, let us build the two datasets as panels.
bd1 = starwars[1:5, c(1, 3)]
bd1 = rbind(bd1, bd1) %>%
mutate(year = c(rep(2021, 5), rep(2022, 5)),
# if the year is not 2021, multiply by a random draw from N(1, 0.025)
mass = ifelse(year == 2021, mass, mass*rnorm(10, 1, 0.025))) %>%
select(name, year, mass) %>%
arrange(name, year)
bd1
## # A tibble: 10 × 3
## name year mass
## <chr> <dbl> <dbl>
## 1 C-3PO 2021 75
## 2 C-3PO 2022 75.5
## 3 Darth Vader 2021 136
## 4 Darth Vader 2022 135.
## 5 Leia Organa 2021 49
## 6 Leia Organa 2022 51.3
## 7 Luke Skywalker 2021 77
## 8 Luke Skywalker 2022 75.4
## 9 R2-D2 2021 32
## 10 R2-D2 2022 33.8
bd2 = starwars[c(2, 4, 7:9), 1:2]
bd2 = rbind(bd2, bd2) %>%
mutate(year = c(rep(2021, 5), rep(2022, 5)),
# if the year is not 2021, height grows by 2%
height = ifelse(year == 2021, height, height*1.02)) %>%
select(name, year, height) %>%
arrange(name, year)
bd2
## # A tibble: 10 × 3
## name year height
## <chr> <dbl> <dbl>
## 1 Beru Whitesun lars 2021 165
## 2 Beru Whitesun lars 2022 168.
## 3 Biggs Darklighter 2021 183
## 4 Biggs Darklighter 2022 187.
## 5 C-3PO 2021 167
## 6 C-3PO 2022 170.
## 7 Darth Vader 2021 202
## 8 Darth Vader 2022 206.
## 9 R5-D4 2021 97
## 10 R5-D4 2022 98.9
- Notice that each character now has two rows, corresponding to the two years, 2021 and 2022. We can therefore run a
full_join()using bothnameandyearas key variables.
# Merge the datasets
full_join(bd1, bd2, by=c("name", "year"))
## # A tibble: 16 × 4
## name year mass height
## <chr> <dbl> <dbl> <dbl>
## 1 C-3PO 2021 75 167
## 2 C-3PO 2022 75.5 170.
## 3 Darth Vader 2021 136 202
## 4 Darth Vader 2022 135. 206.
## 5 Leia Organa 2021 49 NA
## 6 Leia Organa 2022 51.3 NA
## 7 Luke Skywalker 2021 77 NA
## 8 Luke Skywalker 2022 75.4 NA
## 9 R2-D2 2021 32 NA
## 10 R2-D2 2022 33.8 NA
## 11 Beru Whitesun lars 2021 NA 165
## 12 Beru Whitesun lars 2022 NA 168.
## 13 Biggs Darklighter 2021 NA 183
## 14 Biggs Darklighter 2022 NA 187.
## 15 R5-D4 2021 NA 97
## 16 R5-D4 2022 NA 98.9
- Also pay attention to the variable names. When we join datasets that share variables with the same names, but those variables are not used as keys, the function keeps both versions and renames them by default with suffixes
.xand.y. These suffixes can be changed with thesuffixargument.
bd2 = bd2 %>% mutate(mass = rnorm(10)) # create a variable named mass
full_join(bd1, bd2, by=c("name", "year"))
## # A tibble: 16 × 5
## name year mass.x height mass.y
## <chr> <dbl> <dbl> <dbl> <dbl>
## 1 C-3PO 2021 75 167 1.04
## 2 C-3PO 2022 75.5 170. -1.36
## 3 Darth Vader 2021 136 202 -0.329
## 4 Darth Vader 2022 135. 206. 1.07
## 5 Leia Organa 2021 49 NA NA
## 6 Leia Organa 2022 51.3 NA NA
## 7 Luke Skywalker 2021 77 NA NA
## 8 Luke Skywalker 2022 75.4 NA NA
## 9 R2-D2 2021 32 NA NA
## 10 R2-D2 2022 33.8 NA NA
## 11 Beru Whitesun lars 2021 NA 165 0.248
## 12 Beru Whitesun lars 2022 NA 168. -0.581
## 13 Biggs Darklighter 2021 NA 183 -0.613
## 14 Biggs Darklighter 2022 NA 187. 0.286
## 15 R5-D4 2021 NA 97 -0.138
## 16 R5-D4 2022 NA 98.9 0.723